This is the second in a series of blog posts describing updates in the version 2.0 release of gene.iobio. In this post, we discuss the concept of filter status and how to display the variants you want to see. If we open the example data set, click on the 'Filter' button to the left, you will see the 'Filter status' at the top of the left panel.
First of all, what is the filter status anyway? The VCF file was likely generated using variant calling software, for example, Freebayes, then post-processed with a number of tools. One task that may well have been performed is filtering, where a set of filters are set (e.g. quality has to be greater than 1, or the depth at the locus has to be greater than 10 etc.) and each variant is tested against these filters. Rather than throwing away variants that fail filters, the usual practice is to tag the variants with the result of the filtering. If the variant passed, it is often tagged with the value 'PASS'. If not, it is sometimes given a value based on the filter it failed, e.g. 'DEPTH<10'. If the variant failed multiple filters, it can have multiple values separated by a comma, e.g. DEPTH<10,QUAL<1. It is very common to see VCF files where passing variants are given the value 'PASS' and failing variants are not tagged at all. When no tag is given, the VCF field that stores the tag, has the value '.' set for the variant and so this is what we see in gene.iobio. Be careful with the '.' tag though, it doesn't mean fail! It just means no tag has been set, so if no filtering was performed on the VCF file, all variants will have the '.' tag. If 'PASS' is present for some variants and another variant has the filter status '.', it means fail only by the absense of 'PASS'!
As an example, the NA12878 Platinum variant calls for BRCA1 are shown below:
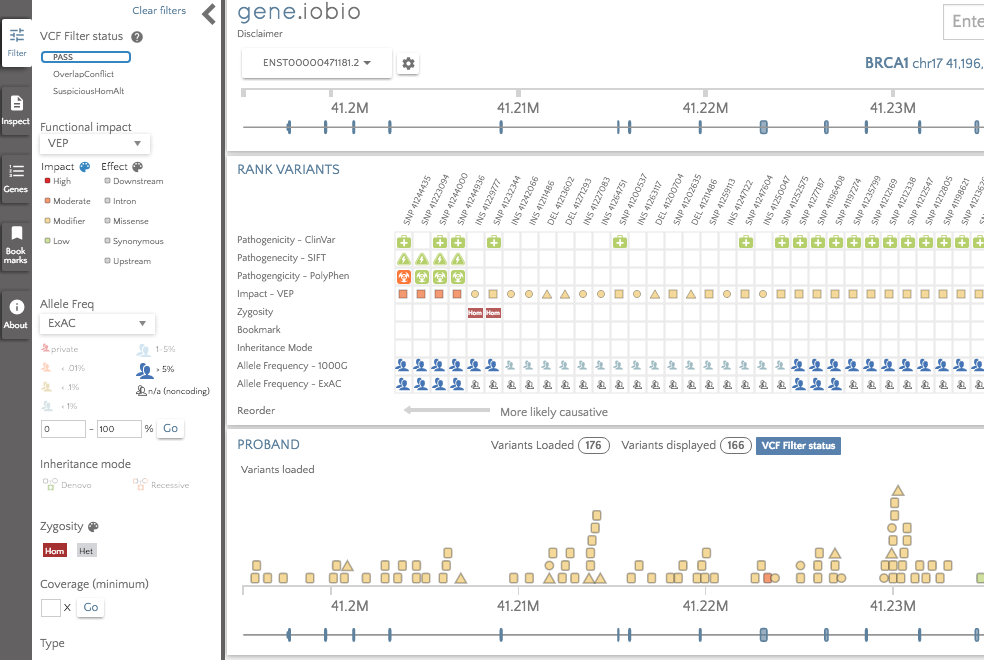
First of all, we can see from the PROBAND variant card that there are 176 loaded variants in this gene, of which only 166 are displayed. The 'VCF Filter status' badge appearing next to these counts indicates that these variants have been removed because of their filter status. If we look in the 'Filter' panel at the left, we can see that there are a number of different tags. At the top is 'PASS' and gene.iobio is, by default, only displaying these variants. The remaining tags, OverlapConflict and SuspiciousHomAlt describe the filters failed by the other variants. If we deselect the 'PASS' variants and select the 'OverlapConflict' variants, we see the variants below:
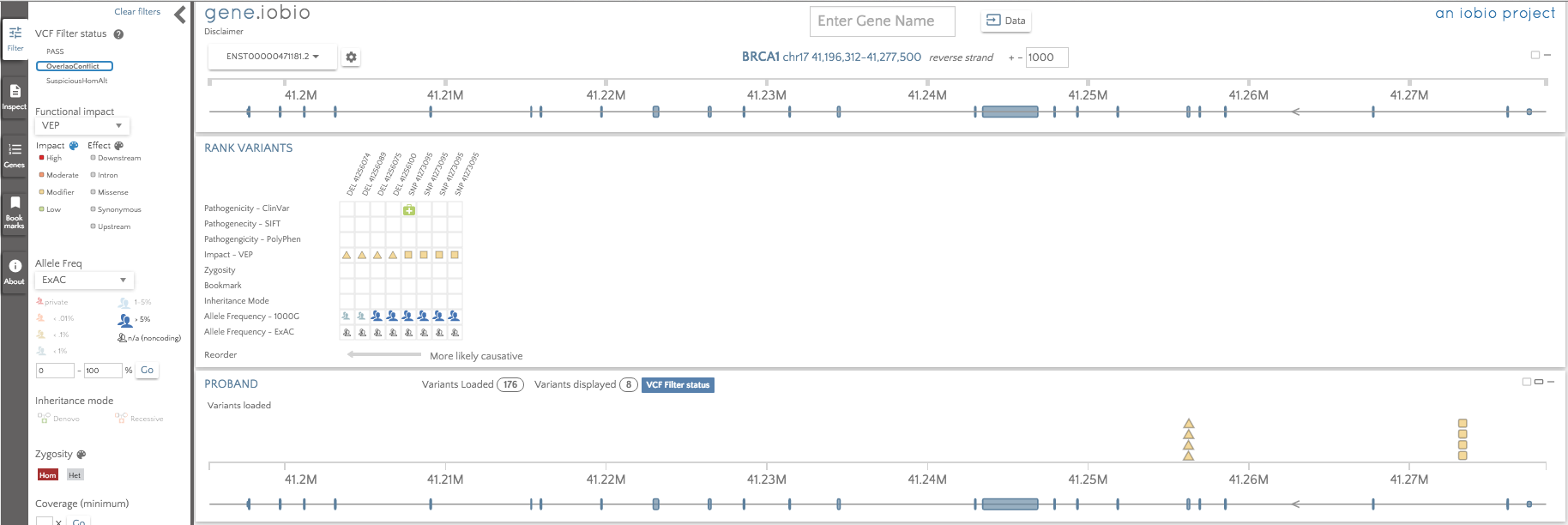
There are four overlapping deletions and four overlapping SNPs. In filtering this file, the software determined that multiple overlapping variants in a single sample is suspicious and tagged them with the OverlapConflict tag to suggest to us, that we should probably not use these variants, or at least use caution.
By displaying only 'PASS' variants by default, gene.iobio omits these suspicious variants, so we don't waste our time analyzing them. But by giving you the option to display them, you can determine for yourself if you think the variants should be filtered out. But, we recommend you analyze your 'PASS' variants first!



