The week, we have pushed out some new features in gene.iobio.
Multi-sample VCF files
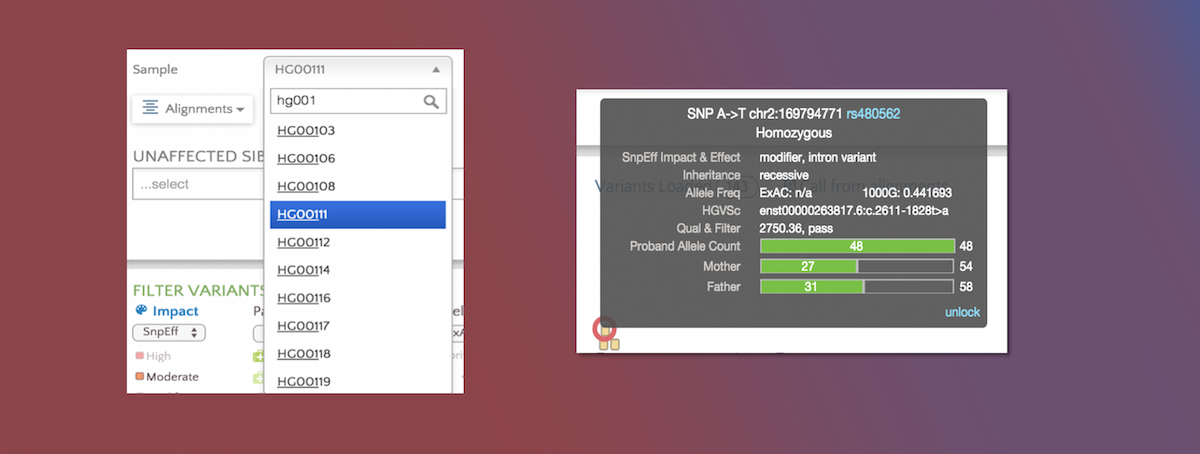
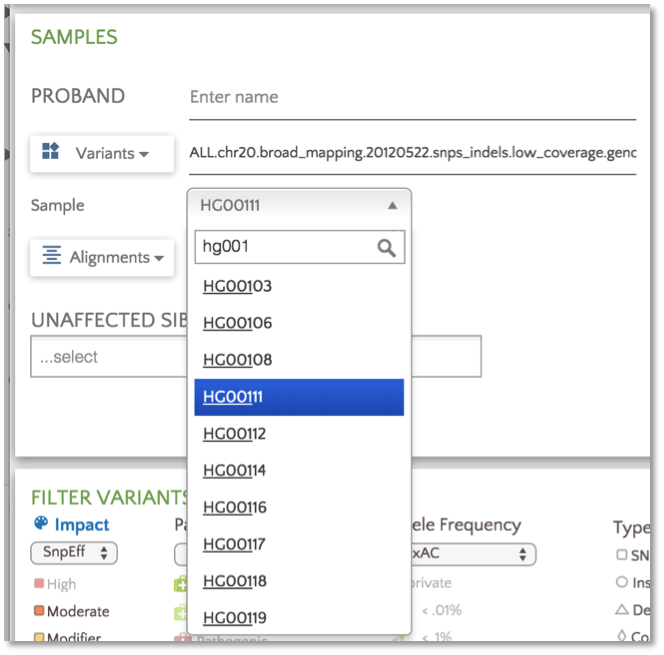
We now support multi-sample VCF files. After entering the VCF URL or selecting the VCF file, a dropdown will appear if the file contains multiple samples.
Here we show a VCF file containing 100+ samples. The sample list can be narrowed down using the type-ahead feature.
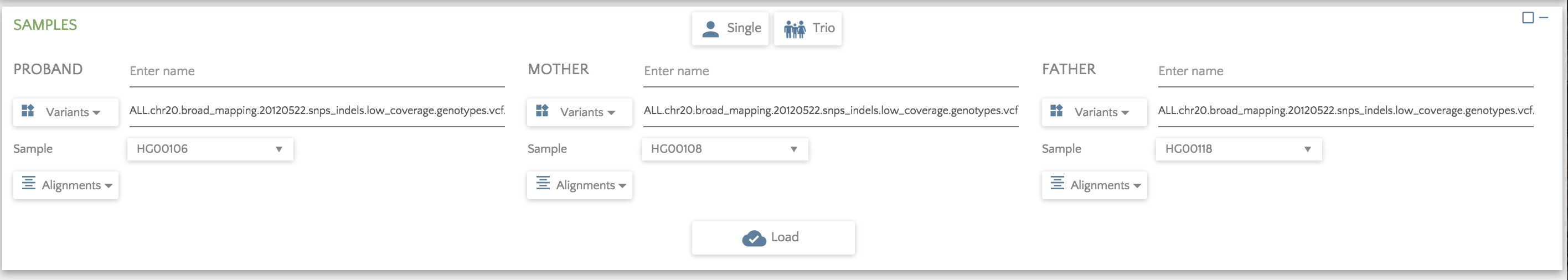
Here is a screen capture showing a trio loaded from one file. Even though the file might be quite large, gene.iobio only examines the samples of interest.
Secure File Serving
Gene.iobio can now handle files served using the https protocol, ensuring secure communication over the network – a requirement when analyzing sensitive data.
Allele Counts
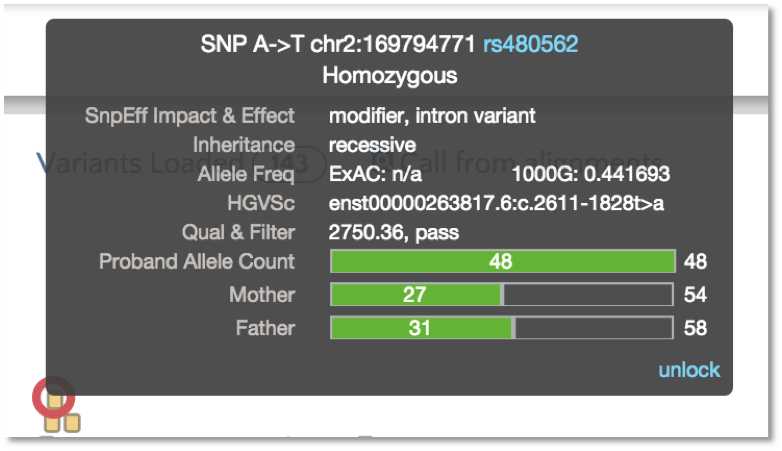
Hover over a variant and the tooltip will include a bar for allele counts. In this example, you can see the allele counts for a trio. Notice how the parents are heterogous alternates and the proband is homozygous alternate. The allele counts verify that there is adequate sequence coverage to make these calls.
Stay tuned. More features are coming soon to gene.iobio including multi-gene navigation and extended trio functionality to analyze variants from unaffected siblings.