Developerα
Alpha Release
Disclaimer
This is an Alpha release. There may be bugs and the api may change moving forward. Thats the downside, the upside is that it's a great time to improve the library with community help and feedback . If you'd like to contribute please feel free to jump in and/or post issues/features on github.
About IOBIO
How it works
At its core iobio is made up of an app and 1 or more web services. To do analysis or data manipulation, input data (via file or url) is passed through 1 or more web services and returned to the app for visualization. This requires somewhat complex websocket code on the app and socket code on the service to correctly analyze the data at each step of the "pipeline" and pass it onto the next webservice. To make all of this simplier we have created several libraries that manage creating the iobio commands/pipelines in the app (iobio.js), creating custom webservices (Minion), and visualizing the streaming data (iobio.viz).
Faster Developement, More Apps
Iobio is built around the philosophy of trying to make genomic web apps easier to build to encourage numerous more focused apps as opposed to big apps that do everything. Towards this goal we've built the back-end as a series of web-services that perform a specific atomic function (e.g. variant calling). These web services can be mixed and matched to support many different web apps. The goals is that when building a new web app, a developer would only need to build the UI/front-end and then simply select the web services they need to support it - no backend required! We are now offering these web services to the community to try to spur even more genomic web apps. Unrestricted the resources for this would be immense, so we have a queue system to ensure that we aren't overloaded with requests. See Queued Web Services to see how this works and how to inform the users where they are in the queue.
Getting Started
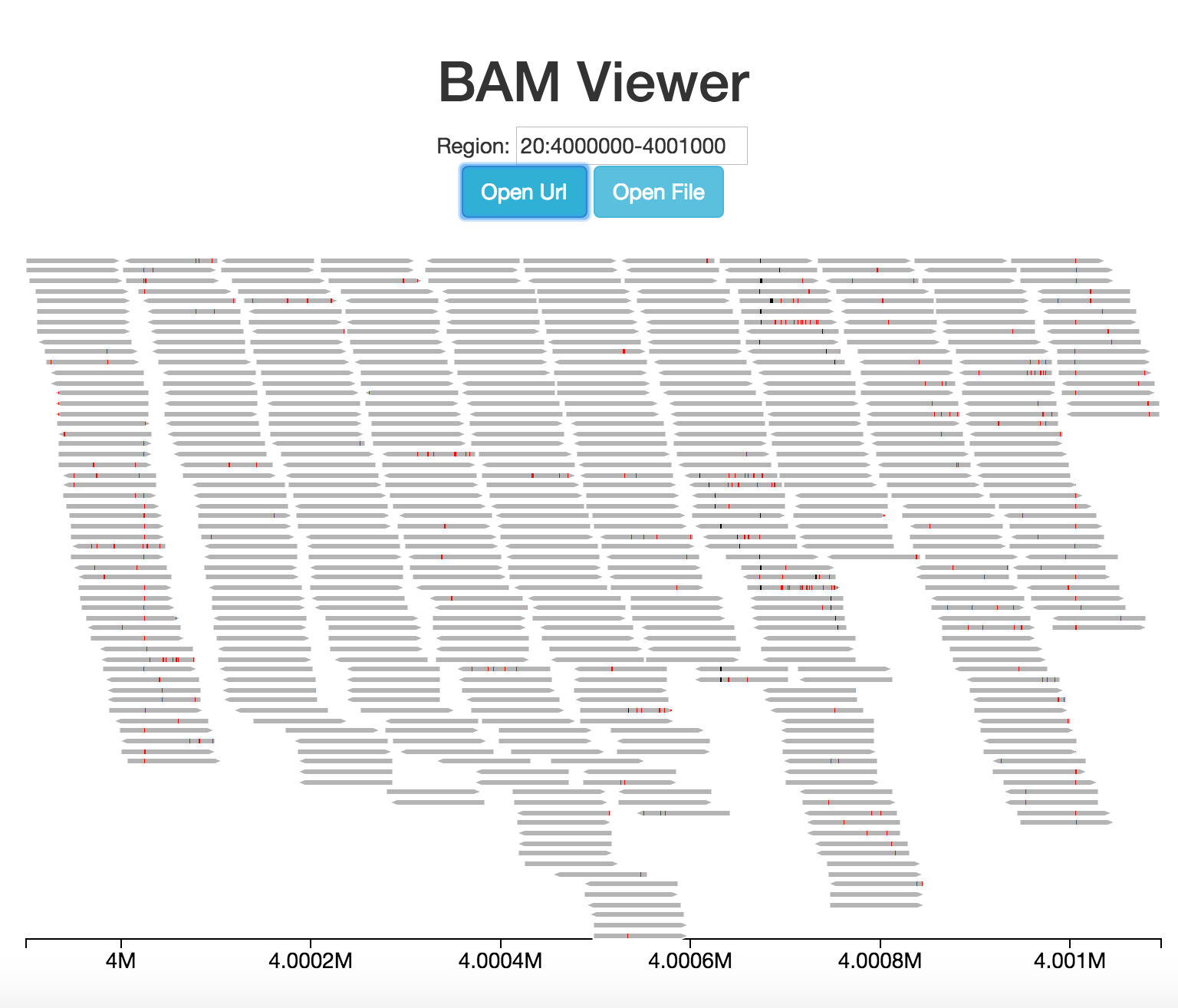
Visualizations
Queued Web Services
Iobio Libraries
The client-side library that handles all the websocket connection code and the construction of iobio commands/pipelines. Full docs and examples here.
The visualization library that is designed to handle streaming data. Full docs and examples here.
The server library that wraps every iobio web service. A goal of the project is to make it easy to create webservices. So Minion enables any command line program that takes data on stdin and generates results on stdout to be converted into an iobio webservice with as little as a 4 line config file. Once wrapped the new iobio webservice can interact with any other iobio webservice in the same was as it could on the command line. Full docs and examples here.